Table of Contents
SOAPdenovo2
Team composition
| Name | |
| Charles Markello | cmarkell@ucsc.edu |
| Thomas Matthew | thjmatth@ucsc.edu |
| Nedda Saremi | nsaremi@ucsc.edu |
SOAPdenovo2 Overview
Short Oligonucleotide Analysis Package de novo (SOAPdenovo) is a novel short-read assembly method that can build a de novo draft assembly for the human-sized genomes. The program is specially designed to assemble Illumina GA short reads.
SOAPdenovo2, the latest version of SOAPdenovo, has the advantage of a new algorithm design that reduces memory consumption in graph construction, resolves more repeat regions in contig assembly, increases coverage and length in scaffold construction, improves gap closing, and optimizes for large genome.
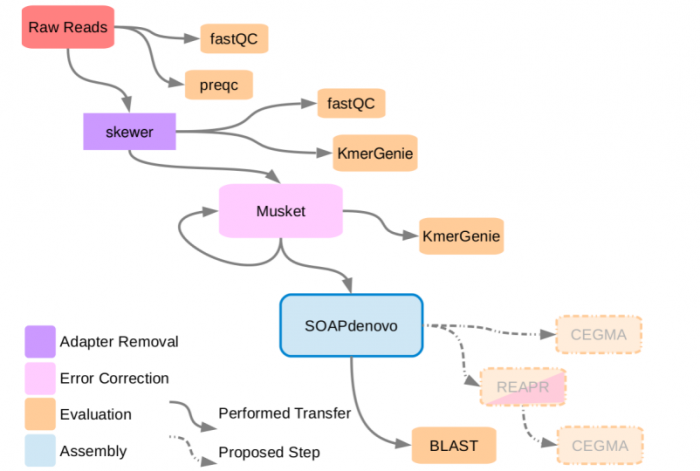
Workflow
Pre-Processing
The following are tools used for processing of the reads prior to assembly using SOAPdenovo2
Skewer
Skewer is an adapter trimming tool specially designed for processing illumina paired-end sequences.
Fastuniq
Fastuniq is a replicates removal tool for de novo assembly
All data sets from the same library were merged and run through Fastuniq
Musket
Musket was used for error correction on the data sets.
KmerGenie
KmerGenie was used to determine optimal k-mer size for de novo assembly. The program produces abundance histograms for many values of k, and then predicts the number of distinct genomic k-mers, returning the k-mer length which maximizes this number.
Running SOAPdenovo2
Configuration File
The configuration file provides the assembler with the information required about each library used for the assembly
Kmer selection
One of the key advantages of SOAPdenovo2 is the ability to select a range of k-mers for the de Bruijn graph assembly step of the genome assembly. This feature is activated through the use of two commands:
-K to select the low end of the kmer range
-M to select the high end of the kmer range
Our Post-Processing
The following are tools used for processing/analysis of the reads after running SOAPdenovo2
CEGMA
Core Eukaryotic Genes Mapping Approach
Builds a highly reliable set of gene annotations in the absence of experimental data. Defines a set of 458 core proteins that are present in a wide range of taxa. Since these proteins are highly conserved, sequence alignment methods can reliably identify their exon-intron structures in genomic sequences.
QUAST
QUality ASsessment Tool
QUAST performs fast and convenient quality evaluation and comparison of genome assemblies.
QUAST computes a number of well-known metrics, including contig accuracy, number of genes discovered, N50, and others
Assembly results
| Attempt | Libraries | File location | Contig/Scaffold/Statistics file | Stats file dump | Run log file |
| 1 | SW018_S1, SW019_S1, SW019_S2 | /campusdata/BME235/S15_assemblies/SOAPdenovo2/assemblyTask/SOAPdenovo2_run1/ | soapdenovo2_sparseGraph.contig, soapdenovo2_sparseGraph.scafSeq, soapdenovo2_sparseGraph.scafStatistics | run1 stats | runlog |
| 2 | SW018_S1, SW019_S1, SW019_S2, R1_IJS8_mates_ICC5_SW023_S60 | /campusdata/BME235/S15_assemblies/SOAPdenovo2/assemblyTask/SOAPdenovo2_run2/ | soapdenovo2_sparseGraph.contig, soapdenovo2_sparseGraph.scafSeq, soapdenovo2_sparseGraph.scafStatistics | run2 stats | runlog2 |
| 3 | SW018_S1, SW019_S1, SW019_S2, R1_IJS8_mates_ICC5_SW023_S60, BS-MK, BS-tag, SW041, SW042, UCSF SW019, UCSF SW018 | /campusdata/BME235/S15_assemblies/SOAPdenovo2/assemblyTask/SOAPdenovo2_run3/ | soapdenovo2_allAssembly_1.contig, soapdenovo2_allAssembly_1.scafSeq, soapdenovo2_allAssembly_1.scafStatistics | run3 stats | runlog3 |
GapCloser
SOAPdenovo2's GapCloser tool was run on the scaffold file to fill in gaps of Ns
Resultant file located here:
/campusdata/BME235/S15_assemblies/SOAPdenovo2/assemblyTask/SOAPdenovo2_run3_all/soapdenovo2_allAssembly_1.scafSeq.gapclosed
QUAST was then used to assess how successful GapCloser was on the scaffold file
GapCloser significantly decreased the number of Ns in the gap-closed scaffold from roughly 17% to 1.2%, resulting in the decrease in the genome size approximation, the Size_includeN parameter
| Parameter | scaffold file | gap closed scaffold file |
| Size_includeN | 2587459468 | 2322694661 |
| Number of Scaffolds | 2427310 | 2427310 |
| Longest_Seq | 124995 | 114336 |
| # N's per 100 kbp | 17265.92 | 1249.37 |
| N50 | 12629 | 11000 |
| L50 | 59060 | 60106 |
QUAST will take inputted scaffolds and break them at repeats of Ns to create theoretical contigs
GapCloser roughly tripled the size of the longest theoretical contig, seen in the longest sequence parameter. It roughly quadrupled the N50 and thus decreased the number of theoretical contigs.
| Parameter | theoretical contig file | theoretical gap closed contig file |
| Size_includeN | 2140718695 | 2293889058 |
| Number of Contigs | 3849281 | 2598123 |
| Longest_Seq | 31799 | 94266 |
| N50 | 1814 | 8033 |
| L50 | 315423 | 80940 |
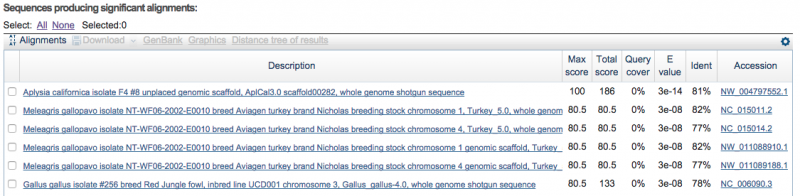
BLAST results
May 26 2015 BLASTN
Searching only NCBI Genomic Reference Sequences
Using the 6 largest scaffolds from the full assembly (Run 3)
Aplysia californica was first when sorted by E value
It was the first mollusk to be sequenced