Table of Contents
Insert Length Analysis (lecture notes)
We discussed the results from mapping the Illumina run 2 reads to the 454 reads using BWA. There is a more complete analysis at Determining Paired-end Insert Size.
Template Length
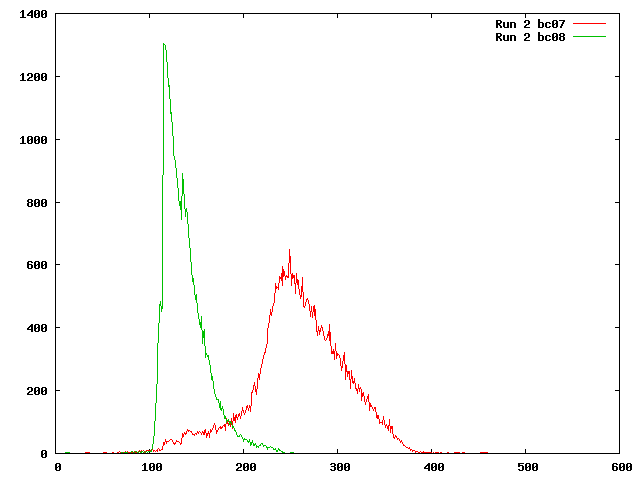
First, we determined what this histogram really shows in the distribution of template lengths, the length from the leftmost base mapped to the rightmost. This size includes the length of each read in the pair and the insert region between them.
Both these distributions show smaller template lengths than the previous estimates (see computer_resources:data). The distribution for barcode 8 is especially odd because it appears to be cut off at 100. The read lengths for both barcodes were around 100 bps so any template length less than 200 represents a pair that can be joined. It was decided that SeqPrep should be run prior to mapping the reads to avoid these pairs that overlap.
454 Coverage
One possible explanation for misshapen distribution is that BWA had difficulty aligning the Illumina reads to the reference due to the possible overlap of the 454 reads. We calculated the expected number of overlaps in the 454 reads.
R = # of 454 reads ( ~500,000 ) G = Length of Genome L = Length of each read ( ~400 ) C = Coverage ( 0.1 for 454 ) = R * L / G The probability of overlapping in one direction is: P_overlap = L * P_read_starts = L * 1/G So for all reads s and t: sum for reads s,t ( L/G ) ~= R^2 * L/G = R * C We expect to see 500,000 * 0.1 = 50,000 reads overlap.
90 percent of our reference does not overlap, so this is not the problem.
Expected Illumina Overlaps with 454
The template lengths we observe come from the Illumina read pairs that map onto the same 454 read. We estimated the expected number of pairs.
L = Length of 454 Read ( ~400-500 ) M = Length of Illumina template ( ~300 for bc07, ~200 for bc08 ) R = # of 454 reads ( ~500,000 ) S = # of Illumina reads ( 1.4e6 for bc07, 11.7e6 for bc08 ) G = genome size The expected number of Illumina templates that map to 454 reads: (L - M) / G * R * S bc07: (200) / (2e9) * (5e5) * (1.4e6) = 7e4 (actual=58,218) bc08: (300) / (2e9) * (5e5) * (11.7e6) = 8e5 (actual=45,324)
We have around the expected number of hits for barcode 7 but much less for barcode 8.