Table of Contents
Overview
BWA is a tool for aligning short reads to a reference genome using two different algorithms based on the Burrows-Wheeler Transform (BWT).
Install
The source for BWA can be found on the BWA SourceForge page. The archive (v0.5.9) was extracted to the /campusdata/BME235/programs directory, compiled and the executable was copied to /campusdata/BME235/bin.
Using BWA
The first step in using BWA is building a reference database.
bwa index -a is database.fasta
Their are two options for the algorithm. The default option, is, is relatively fast and works on genomes smaller than 2GB. The other algorithm, bwtsw, is slower and less accurate but works on longer reads and works with larger databases.
Next, the reads are aligned to the reference using the aln command.
bwa aln database.fasta short_read.fastq > aln_sa.sai
The reads in the fastq file are aligned against the reference data base and the results are written to standard output in the .sai format.
The 'samse' and 'sampe' commands are used to generate single-end and paired-end alignments in SAM format from the FASTQ and SAI files.
bwa samse database.fasta aln_sa.sai short_read.fastq > aln.sam bwa sampe database.fasta aln_sa1.sai aln_sa2.sai read1.fq read2.fq > aln.sam
Note that BWA does seem to accept gzipped files, so there is no need to ungzip the read files, though the documentation doesn't mention this.
Quirks
The SAM formatted alignments include a column labeled “inferred insert length” by the BWA manual, but in the SAM specification it is described as the “template length” or distance between the leftmost mapped base to the rightmost mapped base. The second description seems to match the output of BWA. However, there are some template lengths that do not appear to be calculated correctly.
bc07_1.fastq: @HWUSI-EAS1722:4:66:6286:18215#CAGATC/1 AGCAGTCGTCGTGGTATGCCTGGATGTTACAGCAGTCGTCGTGGTATGACTGGATGTTACAGCAGTCGTCGTGGTATGACTGGATGTTACAGCAGTCGTCGTGGTATGACTGGAT bc07_2.fastq: @HWUSI-EAS1722:4:66:6286:18215#CAGATC/2 CACGACGACTGCTGTAACATCCAGGCATACCACGACGACTGCTGTAACATCCAGGCATACCACGACGACAGCTATAACATACACTCATACCACGA
For example, these two reads make up a pair that overlaps.
...ACATCCAGTCATACCACGACGACTGCTGTAACATCCAGGCATACCACGACGACTGCT
CACGACGACTGCTGTAACATCCAGGCATACCACGACGACTGCTGTAACATCCAGGCATACCACGACGACAGCTATAACATACACTCATACCACGA
Instead of reporting the total length, the length of the overlap is reported.
HWUSI-EAS1722:4:66:6286:18215#CAGATC 81 GAZ7HUX03HIJAL 272 23 115M = 344 -43 HWUSI-EAS1722:4:66:6286:18215#CAGATC 161 GAZ7HUX03HIJAL 344 25 95M = 272 43
This explains the incorrect short lengths found in our histograms. This does not appear to affect the pairs that do not overlap and most of these overlapping reads that should be combined using SeqPrep.
Determining Paired-End Insert Size
BWA was used to estimate the distribution of insert sizes in the Illumina runs for banana slug. The 454 reads were used as the reference and the Illumina reads were mapped onto them. The distribution of the insert lengths can be inferred from the pairs that map onto the same 454 read. This is possible because our insert sizes are smaller than the size of the 454 reads.
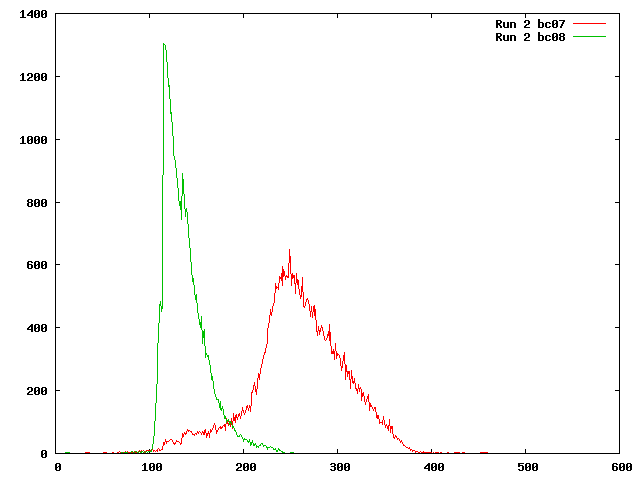
Here is the frequencies of each inferred insert length from the SAM file from the paired end alignments for Illumina run 2. The mean inferred insert size for the barcode 7 reads is 258 bases and 138 bases for the barcode 8 reads. Note that this differs considerably from the estimates of 411 bp for barcode 7 and 372bp for barcode 8 from the computer_resources:data page, which was based on bioanalyzer results for the DNA library. What is the discrepancy? The difference is that the Bioanalyzer results include the adapters, not just the DNA that is sequenced, so the difference is actually fairly small.
Why does the barcode 8 graph cut off so abruptly? (overlapping reads?)
So what exactly is the “inferred insert length” being plotted here? After looking at the SAM format specification and some of the entries in the SAM files, it appears we are actually looking at the “template length”, the total length of each end read plus the insert size.
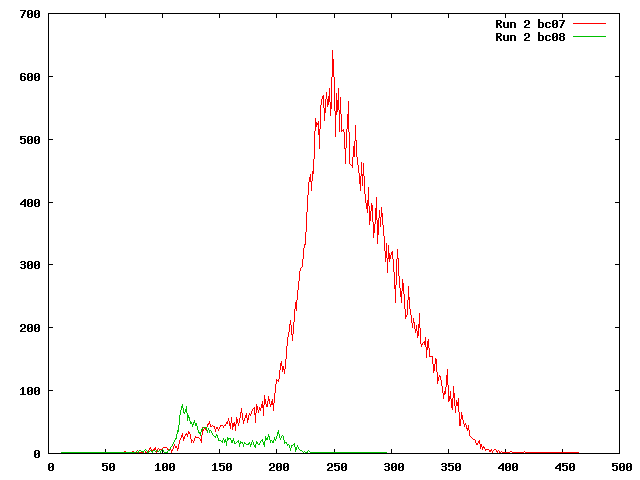
After SeqPrep
We ran SeqPrep on run 2 to remove the Illumina adapter sequences and merge pairs that overlapped and mapped the remaining pairs to the 454 reference. SeqPrep removed most of the barcode 8 pairs that were mapped previously, but left most of the barcode 7 pairs that previously mapped unchanged.
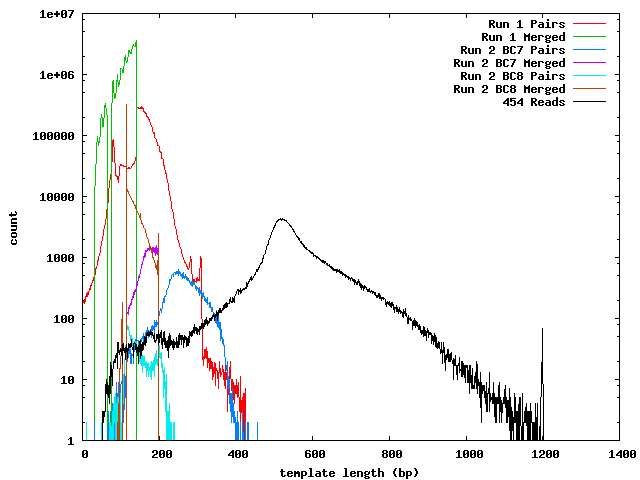
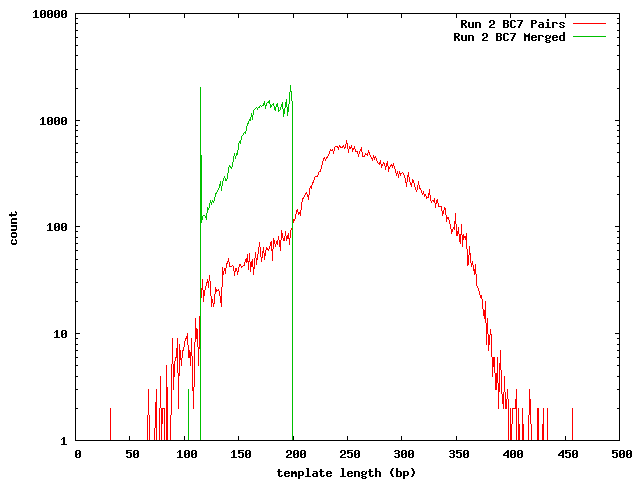
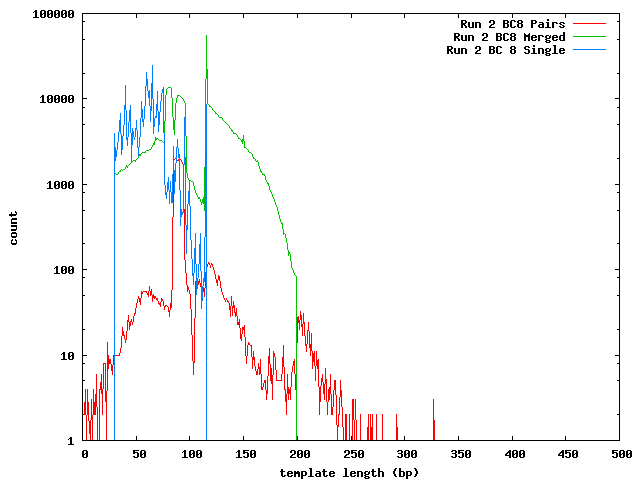
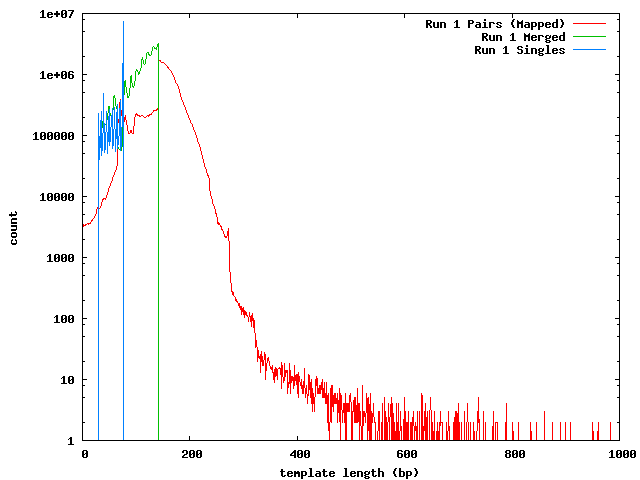
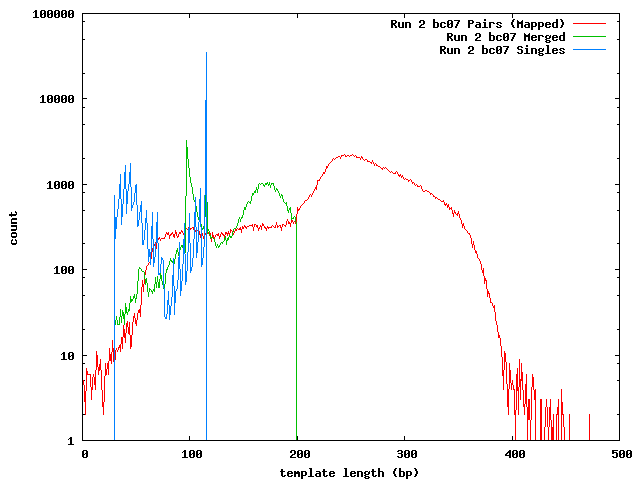
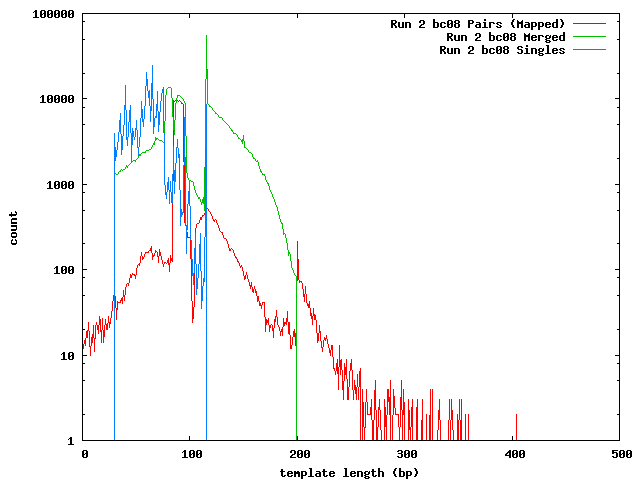
The next histogram shows the 454 length distribution with the SeqPrep merged read lengths and the mapped lengths for the paired-end templates. From this we can see that the lengths of the 454 reads are greater than the illumina templates and the short templates lengths are not due to the lengths of the sequences in the 454 reference.
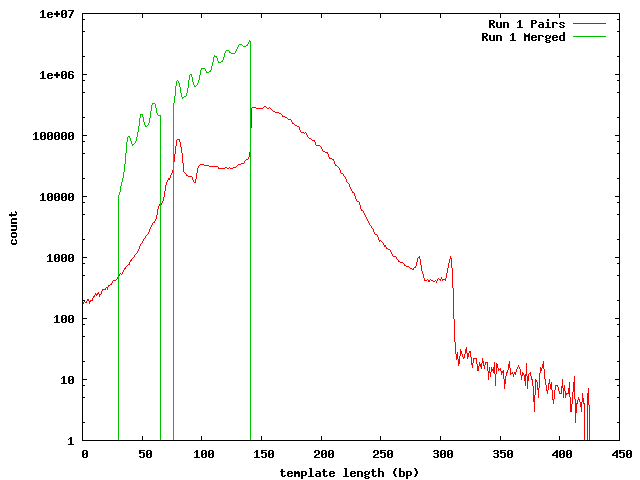
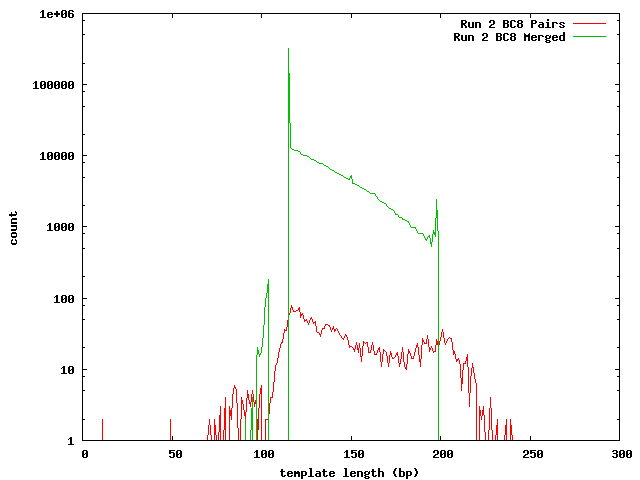
These histograms show the mapped lengths for the paired-end templates and the lengths of merged reads from SeqPrep. In each of these, we can see the distinct range for the SeqPrep merged reads and the split between merged and unmerged pairs. The counts for the paired-end reads are lower than the counts for merged reads because the paired-end reads had to map to one of the 454 reads. We can see that if the paired-end counts were increased by a factor of 10 (for the 0.1x coverage in the 454 reads) the merged and paired-end reads would form a continuous distribution. Lengths less than 90 may be incorrect. The higher frequency of these in run 1 can be explained its higher coverage.
In the merged lengths for both run 1 and run 2 barcode 8 there is a gap of 10 lengths (66-75 for run 1, 105-114 for run 2 bc08) where no reads were observed. This is an artifact of SeqPrep's two methods for merging reads and the 10 base overlap requirement for merging.
After Quake
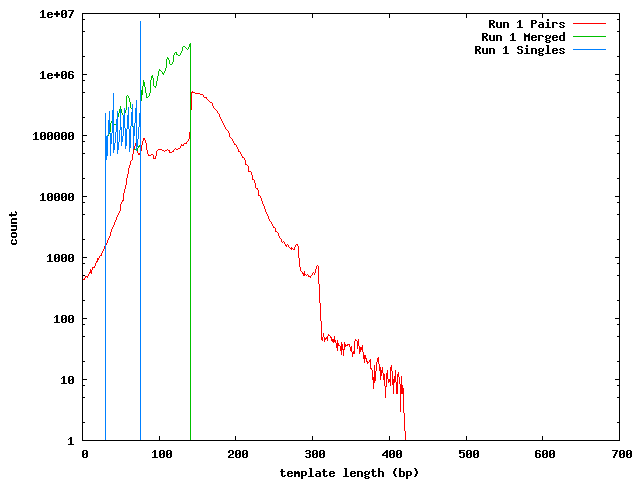
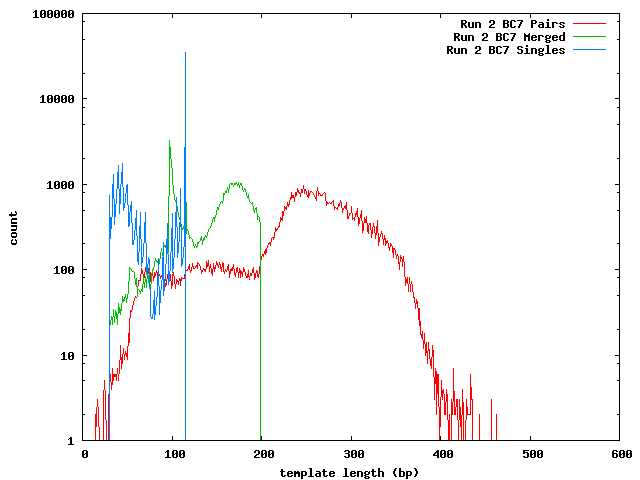
This process was repeated on the data after Quake correction. In addition to the paired reads and merge reads, Quake separates the reads whose pair could not be successfully corrected.
Mean Lengths
Run 1
| Template | 151.7 |
| Merged | 113.3 |
| Single | 65.7 |
Run 2 bc07
| Template | 250.3 |
| Merged | 142.5 |
| Single | 88.8 |
Run 2 bc08
| Template | 94.5 |
| Merged | 104.2 |
| Single | 61.3 |
After Assembly
This process was repeated using the SOAPdenovo contigs from assemblies/slug/SOAPdenovo-assembly2/k47_w_454_contigs instead of the 454 reads as the reference. The shapes of these distributions follow the same patterns as the ones found using the 454 reads, although there are more mapped pairs because of the higher coverage of the contigs and we see some longer templates sizes than before. For the run1 and run2 barcode 7 pairs, the results involve some self-reference, since the previous estimates were used to build the contigs to which we mapped the pairs. However, the pairs for barcode 8 were not used in the assembly because of their negative mean
insert size (most pairs overlap but not in a way that SeqPrep could merge). When these were mapped to the contigs, we saw the same pattern as before but with more reads that mapped successfully.