Table of Contents
Team 4 | ABySS
Assembly By Short Sequences - a de novo, parallel, paired-end sequence assembler
Team composition
| Name | |
| Jared Copher | jcopher@ucsc.edu |
| Emilio Feal | efeal@ucsc.edu |
| Sidra Hussain | sihussai@ucsc.edu |
ABySS overview
ABySS is published by Canada's Michael Smith Genome Sciences Centre, and was the first de novo assembler for large genomes recommended bu Illumina in this technical note when using only their data. The ABySS team are active members on BioStars where they recommend all technical questions be asked.
General notes
- ABySS can run in serial mode, but that isn't too useful for such a large genome.
- The documentation recommends creating assemblies with several values of k and selecting the “best” one.
- The program involves its own error correction.
Installing ABySS
ABySS source code was downloaded from Github
% lftpget https://github.com/bcgsc/abyss/archive/master.zip
ABySS needs to be configured with it's dependencies
% ./autogen.sh % ./configure --prefix=/campusdata/BME235\ % --enable-maxk=96\ #must be a multiple of 32 % --enable-dependency-tracking\ % --with-boost=/campusdata/BME235/include/boost\ % --with-mpi=/opt/openmpi\ % CC=gcc CXX=g++\ % CPPFLAGS=-I/campusdata/BME235/include/
Then ABySS can be installed via the makefile
% make % make install
Notes on Installation
- the -enable-maxk is by default set to 64. Always ensure the max kmer is set to the smallest possible value that accommodates your target kmers so as to minimize ABySS's memory footprint.
- ensure that the CPPFLAGS specified directory holds the google sparsehash includes. Look for a warning in the ABySS compilation to be sure.
- the boost libraries do not need to be compiled before installing.
- ensure openmpi is installed with the –with-sge option (for SGE) to ensure tight integration.
ABySS parameters
Parameters of the driver script, abyss-pe, and their [default value]
- a: maximum number of branches of a bubble [2]
- b: maximum length of a bubble (bp) [10000]
- c: minimum mean k-mer coverage of a unitig [sqrt(median)]
- d: allowable error of a distance estimate (bp) [6]
- e: minimum erosion k-mer coverage [sqrt(median)]
- E: minimum erosion k-mer coverage per strand [1]
- j: number of threads [2]
- k: size of k-mer (bp) [no default]
- l: minimum alignment length of a read (bp) [k]
- m: minimum overlap of two unitigs (bp) [30]
- n: minimum number of pairs required for building contigs [10]
- N: minimum number of pairs required for building scaffolds [n]
- p: minimum sequence identity of a bubble [0.9]
- q: minimum base quality [3]
- s: minimum unitig size required for building contigs (bp) [200]
- S: minimum contig size required for building scaffolds (bp) [s]
- t: minimum tip size (bp) [2k]
- v: use v=-v for verbose logging, v=-vv for extra verbose [disabled]
Please see the abyss-pe manual page for more information on assembly parameters.
Possibly, abyss-pe parameters can have same names as existing environment variables'. The parameters then cannot be used until the environment variables are unset. To detect such occasions, run the command:
abyss-pe env [options]
Above command will report all abyss-pe parameters that are set from various origins. However it will not operate ABySS programs.
Running ABySS
Running an assembly on the campusrocks cluster, which runs on SGE, requires the use of the qsub command. The use of a shell script allows a convenient and concise way to wrap useful qsub options, environmental variable manipulations, and the executable (abyss-pe) itself in a single script. An example script which runs the parallel-version of ABySS on the cluster is shown below.
#!/bin/sh #$ -N team4 #$ -cwd #$ -j y #$ -pe mpi 10 #$ -S /bin/bash #$ -V ABYSSRUNDIR=/campusdata/BME235/bin export PATH=$PATH:/opt/openmpi/bin:/campusdata/BME235/bin/ export LD_LIBRARY_PATH=/opt/openmpi/lib/:$LD_LIBRARY_PATH ABYSSRUN=$ABYSSRUNDIR/abyss-pe $ABYSSRUN np=10 k=21 name=ecoli in='/campusdata/BME235/programs/abyss-1.5.2/JARED/test-data/reads1.fastq /campusdata/BME235/programs/abyss-1.5.2/JARED/test-data/reads2.fastq'
Note that the parallel version of ABySS requires two things in particular:
- The use of a parallel_environment (PE) which can be selected using a qsub option.
- The np option of abyss-pe. The number of processes here must reflect the number included in the parallel environment option.
The PE option in the script above:
#$ -pe mpi 10
The mpi designates the choice of a PE that is installed on the system and the 10 indicates the number of processes over which to run the job. To see which PE's are installed on the system, use the command:
qconf -spl
Selecting the proper PE is critical to ensure the success of a parallelized ABySS job. By using the command:
qconf -sp PE_NAME
you can inspect the settings for an individual PE. For example, using the command qconf -sp mpi will report the following:
pe_name mpi slots 9999 user_lists NONE xuser_lists NONE start_proc_args /opt/gridengine/mpi/startmpi.sh $pe_hostfile stop_proc_args /opt/gridengine/mpi/stopmpi.sh allocation_rule $fill_up control_slaves FALSE job_is_first_task TRUE urgency_slots min accounting_summary TRUE
For using ABySS with openmpi, there are three settings in particular which should be noted:
- slots: indicates maximum number of slots that can be designated with the PE
- allocation_rule: indicates the form of scheduling to be used to allocate slots to nodes
- control_slaves: used to indicate a tightly managed interface
For all of the PE's installed on the campus cluster, the slots were all set at 9999 (virtually limitless) and so were not of terrible concern to us. However, the allocation_rule and control_slaves were critical. The initial PE tried was mpi however, because openmpi can achieve tight integration with the cluster's SGE, we switched to orte. This was found to be inadequate due to its allocation_rule of $pe_slots. This demanded that all slots requested by the job be accommodated on a single node. This proved to be fatal since the memory requirements of a run with full libraries required more memory than that possessed by the largest node (~256 GB). We then switched to mpich which offers both tight integration and an allocation_rule of $fill_up which, when used with a sufficiently large number of slots, would ensure that multiple nodes are used in a job. However, the limitation in this setting is that, depending on the relative traffic on the cluster, we would have to designate more slots than needed to ensure the use of multiple nodes. To address this, we had the denovo PE created and added to the queue. This PE offers both tight integration and an allocation_rule of $round_robin. This allocation setting ensures that the slots requested are evenly distributed across available nodes. Thus, selecting relatively few processes would ensure the use of multiple nodes' memory.
abyss-pe is a driver script implemented as a Makefile. Any option of make may be used with abyss-pe. Particularly useful options are:
-C dir, --directory=dir
Change to the directory dir and store the results there.
-n, --dry-run
Print the commands that would be executed, but do not execute them.
Commands of abyss-pe
- default: Equivalent to `scaffolds scaffolds-dot stats'.
- unitigs: Assemble unitigs.
- unitigs-dot: Output the unitig overlap graph.
- pe-sam: Map paired-end reads to the unitigs and output a SAM file.
- pe-bam: Map paired-end reads to the unitigs and output a BAM file.
- pe-index: Generate an index of the unitigs used by abyss-map.
- contigs: Assemble contigs.
- contigs-dot: Output the contig overlap graph.
- mp-sam: Map mate-pair reads to the contigs and output a SAM file.
- mp-bam: Map mate-pair reads to the contigs and output a BAM file.
- mp-index: Generate an index of the contigs used by abyss-map.
- scaffolds: Assemble scaffolds.
- scaffolds-dot: Output the scaffold overlap graph.
- stats: Display assembly contiguity statistics.
- clean: Remove intermediate files.
- version: Display the version of abyss-pe.
- versions: Display the versions of all programs used by abyss-pe.
- help: Display a helpful message.
Programs in pipeline
abyss-pe uses the following programs, which must be found in your PATH:
- ABYSS: de Bruijn graph assembler
- ABYSS-P: parallel (MPI) de Bruijn graph assembler
- AdjList: find overlapping sequences
- DistanceEst: estimate the distance between sequences
- MergeContigs: merge sequences
- MergePaths: merge overlapping paths
- Overlap: find overlapping sequences using paired-end reads
- PathConsensus: find a consensus sequence of ambiguous paths
- PathOverlap: find overlapping paths
- PopBubbles: remove bubbles from the sequence overlap graph
- SimpleGraph: find paths through the overlap graph
- abyss-fac: calculate assembly contiguity statistics
- abyss-filtergraph: remove shim contigs from the overlap graph
- abyss-fixmate: fill the paired-end fields of SAM alignments
- abyss-map: map reads to a reference sequence (BW transform)
- abyss-scaffold: scaffold contigs using distance estimates
- abyss-todot: convert graph formats and merge graphs
New to Version 1.3.5 (Mar 05, 2013)
- abyss-mergepairs: Merges overlapping read pairs.
- abyss-layout: Layout contigs using the sequence overlap graph.
- abyss-samtobreak: Calculate contig and scaffold contiguity and correctness metrics.
New to Version 1.5.2 (Jul 10, 2014)
- konnector: fill the gaps between paired-end reads by building a Bloom filter de Bruijn graph and searching for paths between paired-end reads within the graph
- abyss-bloom: construct reusable bloom filter files for input to Konnector
ABySS pipeline
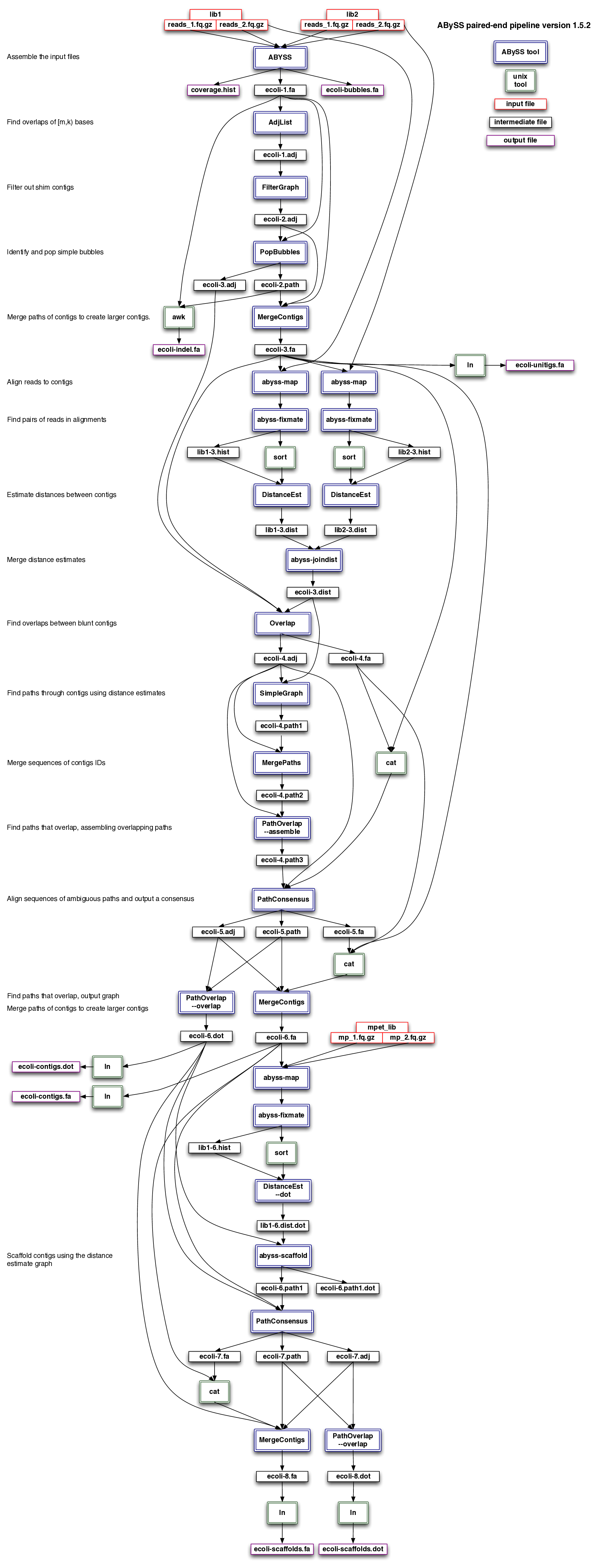
Test run
This run was done using version 1.5.2. The assembly used k=59, 10 processes, and requested mem_free=15g from qsub. The assembly was done using the SW018 and SW019 libraries only. Specifically, the files used were:
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep/SW018_S1_L007_R1_001_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep/SW018_S1_L007_R2_001_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep/SW019_S1_L001_R1_001_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep/SW019_S1_L001_R2_001_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep/SW019_S2_L008_R1_001_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep/SW019_S2_L008_R2_001_trimmed.fastq.gz
The files had adapters trimmed using SeqPrep (see the data pages for more details). SW019_S1 and SW019_S2 were treated as two separate libraries.
The output and log files for this assembly are in /campusdata/BME235/S15_assemblies/abyss/sidra/test_run/singleK.
Results
Note: the N50, etc., stats only include contigs >= 500 bp (I believe the rest are discarded).
There are 10.23 * 10^6 contigs. The N50 contig size is 2,669. The number of contigs of at least N50 (n:N50) is 174,507. The maximum contig size is 31,605, and the total number of bp (in contigs >= 500 bp) is 1.557 * 10^9.
Here are the stats summarized for the contigs and also for scaffolds and unitigs. n:500 is the number of contigs/unitigs/scaffolds at least as long as 500 bp. sum is the number of bases in all the contigs/unitigs/scaffolds at least as long as 500 bp combined.
| n | n:500 | n:N50 | min | N80 | N50 | N20 | E-size | max | sum | name |
| 11.95e6 | 993409 | 247109 | 500 | 962 | 1795 | 3327 | 2296 | 30520 | 1.456e9 | slug-unitigs.fa |
| 10.23e6 | 785054 | 174507 | 500 | 1320 | 2669 | 5079 | 3433 | 31605 | 1.557e9 | slug-contigs.fa |
| 10.11e6 | 711022 | 153036 | 500 | 1490 | 3063 | 5870 | 3945 | 37466 | 1.573e9 | slug-scaffolds.fa |
Notes
The success of this run means we are probably ready to do a run with all the data (not including the mate-pair data, that can be used for scaffolding later). For that run, the different trimmed files for each library should be concatenated, so that the run involves only the actual number of libraries we had (I believe 4?). It should also use many more than 10 processes.
Test with more data
This run was also done using version 1.5.2. I used SGE job arrays to submit multiple assemblies with different k values, k=55-65 and 27-37, odds only. (Using this many similar ks is probably unnecessary.) The syntax for submitting an array w/ multiple ks is
qsub -t 55-65:2 slug-pe.sh
where slug-pe.sh uses $SGE_TASK_ID in place of the k value. I used 32 processes per job and tried submitting with various amounts of memory requested, from 10-30g. I also added a quality cutoff of 20 (parameter q=20) to this run. The files used this time were concatenated versions of the various sequencing runs for each library:
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R1_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R2_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019_S1_S2_and_UCSF_R1_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019_S1_S2_and_UCSF_R2_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-MK_CATCCGG_R1_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-MK_CATCCGG_R2_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R1_trimmed.fastq.gz
- /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R2_trimmed.fastq.gz
The files had adapters trimmed using SeqPrep, but were not merged (see the data pages for more details). The four libraries used were SW018, SW019, BS-MK, and BS-tag.
I was not able to get these to run successfully. On the first attempt, the job with k=55 started running and died after 6 hours. It was difficult to tell why based on the error (probably ran out of memory) so after this I added the v=-v (verbose) option. Here's the error I got (before verbose option):
/opt/openmpi/bin/mpirun -np 32 ABYSS-P -k55 -q20 --coverage-hist=coverage.hist -s slug-bubbles.fa - o slug-1.fa /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R 1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and _UCSF_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019 _S1_S2_and_UCSF_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_ libs/SW019_S1_S2_and_UCSF_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPr ep_newData/UCSF_BS-MK_CATCCGG_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/S eqPrep_newData/UCSF_BS-MK_CATCCGG_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimmi ng/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_t rimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R2_trimmed.fastq.gz ABySS 1.5.2 ABYSS-P -k55 -q20 --coverage-hist=coverage.hist -s slug-bubbles.fa -o slug-1.fa /campusdata/BME235/Spr ing2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R1_trimmed.fastq.gz /campusdata/BME 235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019_S1_S2_and_UCSF_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019_S1_S2_and_UCSF_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-MK_CATCCGG_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-MK_CATCCGG_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R2_trimmed.fastq.gz Running on 32 processors 9: Running on host campusrocks2-0-2.local 25: Running on host campusrocks2-0-2.local 2: Running on host campusrocks2-0-2.local 3: Running on host campusrocks2-0-2.local 8: Running on host campusrocks2-0-2.local 10: Running on host campusrocks2-0-2.local 11: Running on host campusrocks2-0-2.local 13: Running on host campusrocks2-0-2.local 27: Running on host campusrocks2-0-2.local 26: Running on host campusrocks2-0-2.local 7: Running on host campusrocks2-0-2.local 12: Running on host campusrocks2-0-2.local 28: Running on host campusrocks2-0-2.local 14: Running on host campusrocks2-0-2.local 18: Running on host campusrocks2-0-2.local 6: Running on host campusrocks2-0-2.local 22: Running on host campusrocks2-0-2.local 30: Running on host campusrocks2-0-2.local 23: Running on host campusrocks2-0-2.local 24: Running on host campusrocks2-0-2.local 1: Running on host campusrocks2-0-2.local 17: Running on host campusrocks2-0-2.local 19: Running on host campusrocks2-0-2.local 0: Running on host campusrocks2-0-2.local 16: Running on host campusrocks2-0-2.local 4: Running on host campusrocks2-0-2.local 15: Running on host campusrocks2-0-2.local 29: Running on host campusrocks2-0-2.local 31: Running on host campusrocks2-0-2.local 5: Running on host campusrocks2-0-2.local 20: Running on host campusrocks2-0-2.local 21: Running on host campusrocks2-0-2.local 0: Reading `/campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R1_trimmed.fastq.gz'... 2: Reading `/campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019_S1_S2_and_UCSF_R1_trimmed.fastq.gz'... 3: Reading `/campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019_S1_S2_and_UCSF_R2_trimmed.fastq.gz'... 7: Reading `/campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R2_trimmed.fastq.gz'... 4: Reading `/campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-MK_CATCCGG_R1_trimmed.fastq.gz'... 1: Reading `/campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R2_trimmed.fastq.gz'... 5: Reading `/campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-MK_CATCCGG_R2_trimmed.fastq.gz'... 6: Reading `/campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R1_trimmed.fastq.gz'... -------------------------------------------------------------------------- mpirun noticed that process rank 0 with PID 8806 on node campusrocks2-0-2.local exited on signal 9 (Killed). -------------------------------------------------------------------------- make: *** [slug-1.fa] Error 137
The job ran for about 6 hours before being killed, according to qacct.
For all other jobs in the array, and for other tries using the same submission script, the jobs never started running, and I got this error:
/opt/openmpi/bin/mpirun -np 32 ABYSS-P -k57 -q20 --coverage-hist=coverage.hist -s slug-bubbles.fa -o slug-1.fa /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW018_S1_and_UCSF_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019_S1_S2_and_UCSF_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_catted_libs/SW019_S1_S2_and_UCSF_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-MK_CATCCGG_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-MK_CATCCGG_R2_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R1_trimmed.fastq.gz /campusdata/BME235/Spring2015Data/adapter_trimming/SeqPrep_newData/UCSF_BS-tag_GTAGAGG_R2_trimmed.fastq.gz
error: executing task of job 81112 failed: execution daemon on host "campusrocks2-0-1.local" didn't accept task
--------------------------------------------------------------------------
A daemon (pid 43664) died unexpectedly with status 1 while attempting
to launch so we are aborting.
There may be more information reported by the environment (see above).
This may be because the daemon was unable to find all the needed shared
libraries on the remote node. You may set your LD_LIBRARY_PATH to have the
location of the shared libraries on the remote nodes and this will
automatically be forwarded to the remote nodes.
--------------------------------------------------------------------------
--------------------------------------------------------------------------
mpirun noticed that the job aborted, but has no info as to the process
that caused that situation.
--------------------------------------------------------------------------
--------------------------------------------------------------------------
mpirun was unable to cleanly terminate the daemons on the nodes shown
below. Additional manual cleanup may be required - please refer to
the "orte-clean" tool for assistance.
--------------------------------------------------------------------------
campusrocks2-0-1.local - daemon did not report back when launched
campusrocks2-0-2.local - daemon did not report back when launched
make: *** [slug-1.fa] Error 1
It seems like the ABySS group in a previous iteration of the class had the same issue. I submitted an IT ticket and received absolutely no help. The only thing I could figure out is that this is some sort of issue with openmpi, so our next steps will be attempting the assembly using orte instead of openmpi and also trying the assembly on edser2. We are also going to try some end trimming first so that we have a better chance of not using up all the available memory.
The output and log files for this assembly are in /campusdata/BME235/S15_assemblies/abyss/sidra/assembly and /campusdata/BME235/S15_assemblies/abyss/sidra/assembly/failed_runs.

