Table of Contents
Quake
Quake is a sequencing error correction program that uses kmer count distributions to make the distinction between trusted (suspected true) and untrusted (suspected erroneous) reads.[1]
Running Quake
To run Quake with the Jellyfish counts, it seems we will want to first use stats to dump the kmers as tab-separated and column (-t and -c respectively):
jellyfish stats -t -c -o [counts file]
Then, build the covariance model to find the cutoff, default is 200:1 likelihood ratio of untrusted to trusted reads (same as plotting the histogram from Jellyfish):
cov_model.py [counts file]
Finally, correct the reads:
correct -f [fastq file list] -k [k-mer size] -c [cutoff] -m [counts file] -p [number of cores] -z (gzips the output)
In the file list, you should tab-separate paired end reads. Also, be sure that all .'s in the sequence are written as N's. If the quality scores are written as Phred + 64, you can use -q 64 to handle it.
K-mer counts can also be pre-filtered to save space.
Quake dev: Once you've decided on a cutoff, Quake ignores all of the k-mers below that cutoff. So sure, you can filter the file to save some disk space. But having all of the k-mer counts is best for choosing the cutoff. My cov_model.py script to automatically choose the cutoff requires them.
[Kevin] I think that we want to select the k-mer size manually, rather than relying on quake. Their default cutoff is very conservative, and we'll probably do better over-correcting than under-correcting. If we look at the hugely over-represented k-mers (like the adapter sequences), and compare the true sequences to one that are one base different, we see that the true ones are about 30 times as frequent. Thus quake's idea of correcting only the rarely seen k-mers isn't quite right. What we really want to correct are those k-mers that are close neighbors of much more frequent k-mers. I've not figured out yet precisely what “much more frequent” should mean.
Potential Problems
- Input files need to have an extension, or Quake will throw a substr error when trying to merge hidden files into a result.
- With paired-end input, Quake will output two files for each paired-end read. One will be the cor.fastq file, which contains corrected, paired reads. The other will be the cor_single.fastq file, which contains reads where only one pair could be corrected. You can treat the cor_single.fastq file as a single read file.
Methods
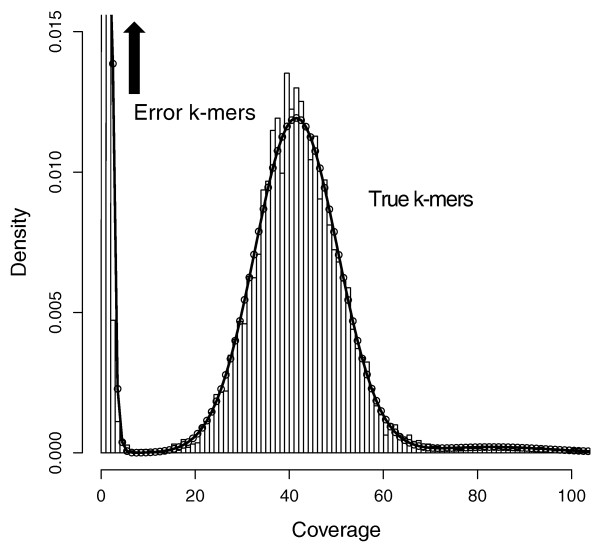
The paper gives an example of the distribution typically seen in kmer counting below. They fit a Gamma distribution to the untrusted reads, and a Gaussian + Zeta (for the high frequency repeats) mixture for trusted reads. The distribution of the trusted reads is actually expected to be Poisson, but the variance is significantly larger than the mean due to sequencing biases. Their example is a high-coverage run, and they note that the distributions will overlap–as we've seen–with lower coverages. This of course makes finding the cutoff trickier. They chose a point where there was a high likelihood ratio of untrusted to trusted read, around 1000:1.
They also recommend a kmer probability of 0.01 in a random sequence that is as long as the genome. That is, 2*G/4^k ~ 0.01, where G is the size of the sequenced genome and k is the size of the kmer. Simplified, k ~ log4(200*G), which is about 19 for our prediction of the banana slug genome size, 2.042e+09.